General Function Checklist
See this page in the users guide: Personal Synonyms
Chromosome synonyms
Here is an example chromosome synonyms file: chromosome.txt
KlingonChr1 Chr1
KlingonChr2 Chr2
KlingonChr3 Chr3
KlingonChr4 Chr4
KlingonChr5 Chr5
Here is the head of an example bed file, made by taking segments from the A_thaliana_Jun_2009 gene annotations and replacing the chromosome names: KlingonGenes.bed
NovelChrom 3630 5899 AT1G01010.1 0 + 3759 5630 0 6 283,281,120,390,153,461, 0,365,855,1075,1543,1808,
KlingonChr1 5927 8737 AT1G01020.1 0 - 6914 8666 0 10 336,633,76,67,86,74,46,90,48,167, 0,509,1229,1456,1636,1834,2014,2308,2489,2643,
KlingonChr1 6789 8737 AT1G01020.2 0 - 7314 8666 0 8 280,294,86,74,46,90,48,167, 0,367,774,972,1152,1446,1627,1781,
KlingonChr1 11648 13714 AT1G01030.1 0 - 11863 12940 0 2 1525,380, 0,1686,
KlingonChr1 23145 31227 AT1G01040.1 0 + 23518 31079 0 20
Notice that there is one entry for a chromosome (NovelChrom) that is not in the synonyms file.
- Navigate to the Data Sources tab in Preferences.
- Click ... next to the Chromosome Synonyms File field.
- Add the chromosome.txt file.
- Restart IGB.
- Open the A_thaliana_Jun_2009 genome.
- Load the KlingonGenes.bed file from the Available Data section.
- Ensure the Load Mode in the Data Access tab at the bottom of IGB is set to Genome.
Good.
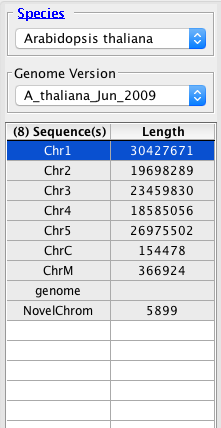
Bad.
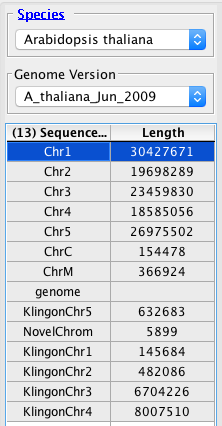
- The annotations from KlingonGenes.bed appear on the same chromosomes as the automatically loaded annotations. (If IGB creates new chromosomes labeled "KlingonChr1" etc, then the link between chromosome "KlingonChr5" and "Chr5" is not functioning.)
- mac
- linux
- windows
Genome version synonyms
Review the user guide page: Use synonyms.txt to link genome version names to each other.
How to make the test file (These instructions are a work in progress)
You can download the test file I made: myQL
Unzip that folder, and continue to How to use the test file.
The following instructions should be viewed as a "fall back".
- Make an empty folder called "myQL".
- In the myQL folder, create a plain text file called contents.txt file with this line (it is a tab between 2018 and My; do not include the bullet):
- A_new_Jun_2018 My new genome as a test
- In myQL, create a folder called "A_new_Jun_2018"
- In A_new_Jun_2018, create an annots.xml file with this text:
<files>
<file name="KlingonGenes.bed" title="new/KlingonGenes" />
</files>
- In the A_new_Jun_2018 folder, save the KlingonGenes.bed file (see Chromosome synonyms).
- In the myQL folder, create a folder called "synsFileNotInUse"
- In the synsFileNotInUse folder, create a text file called synonyms.txt with this text (this is two terms, separated by a tab):
- A_thaliana_Jun_2009 A_new_Jun_2018
The synonyms file will not "work" from this location. This is just its starting position in the test process.
How to use the test file
- Download the myQL test file here: myQL
- Reset preferences to give IGB a clean slate.
- From the IGB home screen, click the Species pull down.
- There is NO genome for A_new in the Species pull down.
- mac
- linux
- windows
- In the Data Sources tab in Preferences, add myQL as a Quickload site.
- From the IGB home screen, click the Species pull down.
- There IS a genome for A_new in the Species pull down.
- mac
- linux
- windows
Open A_new_Jun_2018.
- The myQL Quickload appears in the Available Data section.
- mac
- linux
- windows
- Add KlingonGenes from the myQL Quickload.
- Click Load Data.
- The KlingonGenes data loaded and Klingon chromosomes are now listed in the panel at the right.
- mac
- linux
- windows
Open A_thaliana_Jun_2009.
- There is NO myQL Quickload in the Available Data section.
- mac
- linux
- windows
In the Data Sources tab in Preferences, add synonyms.txt file (found in myQL > synsFileNotInUse) as a version synonyms file.
- IGB prompted you to restart.
- mac
- linux
- windows
Restart IGB.
- There is NO genome for A_new in the Species pull down.
- mac
- linux
- windows
Open A_thaliana_Jun_2009.
- The myQL Quickload appears in the Available Data section.
- mac
- linux
- windows
- Add KlingonGenes from the myQL Quickload.
- Click Load Data.
- The KlingonGenes data loaded and Klingon chromosomes are now listed in the panel at the right.
- mac
- linux
- windows
- Reset preferences to default.
- In your File Chooser, move the synonyms.txt file from the synsFileNotInUse folder to the myQL folder.
- Restart IGB.
- In the Data Sources tab in Preferences, add myQL as a Quickload site.
- There is NO genome for A_new in the Species pull down.
- mac
- linux
- windows
Open A_thaliana_Jun_2009.
- The myQL Quickload appears in the Available Data section.
- mac
- linux
- windows
- Add KlingonGenes from the myQL Quickload.
- Click Load Data.
- The myQL Quickload and the KlingonGenes file were loaded without issue with the A_thaliana_Jun_2009 genome.
- mac
- linux
- windows