Introduction
Graphs contain numerical data associated with base pair positions along the chromosome sequence axis, such as sets of scores or chromosome copy numbers at specific positions. IGB allows you to visualize such graph data and compare it with RefSeq and other annotations.
Examples of suitable data include:
- Files generated from Affymetrix software tools, such as the Expression Console, CNAT, GCOS, or ExACT.
- Expression values from genome tiling arrays.
- Density of ESTs across chromosomes
- GC content along chromosomes
- Measures of conservation between two genomes
There are two basic types of graphs:
- Position graphs associate scores with single genomic positions.
- Interval graphs associate scores with ranges of genomic positions.
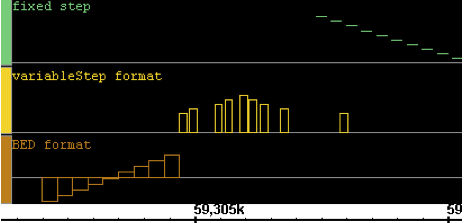
Here is a very simple example of several interval graphs:
Loading graph files
Graph files are loaded in the same way as annotation data files. To open graph files, see instructions for Loading data.
IGB can display graphs in several file formats, developed at Affymetrix or elsewhere. All file names must include the file name extension, such as '.sgr'. Compressed files must include the graph type extension AND the compression extension, such as "mygraph.sgr.zip". For the list of all file formats IGB supports, see File Formats.
Graph files in some formats (notably .gr) do not contain information about which genomes and chromosomes they correspond to, so be sure to set Species and Genome Version before or as part of loading these files. These may be loaded and displayed alongside annotations on any chromosome, but the data may have no meaning if compared to the a chromosome it was not designed for. i.e., to use a .gr file, you must first load the proper organism, genome AND chromosome prior to loading the .gr file
Hiding and deleting graphs
Hide graphs to remove them from the display without deleting them from memory. Right click on the graph track label and choose Hide from the menu, just as you would do for annotation tracks.
Floating graphs must be 'docked' by deselecting Float before they can be hidden.
Hidden graphs can be restored by right clicking in ANY track label and using the Show function. To hide one or more tracks, select the graph tracks or use the Select All button in the Graph Adjuster panel, right click and select Hide. Alternatively, use the Data Management Table to hide and show.
Delete graphs to permanently remove them from the viewer and from memory. To delete graphs, select and click the Delete Graph button in Graphs tab, use the Toolbar shortcut Delete Selected Track or use the red X icon in the Data Access > Data Management Table.
Saving graph tracks
IGB allows you to save individual graph tracks for later use or sharing. This is especially important for graph tracks made in IGB, either through manipulation of your graphs or after using one of IGB's create graph functions. See Saving and Sharing Data.