...
- Observation: The table lists several Quickload sites. (They may be different from those shown below.) If any are shown with a red background, that means that there is a problem with the site - IGB can't reach it. This is a bug and should be reported.
- Ensure that the edit button in the Data Sources window opens a new window and allows a user to modify an app repository URL or to choose a local folder.
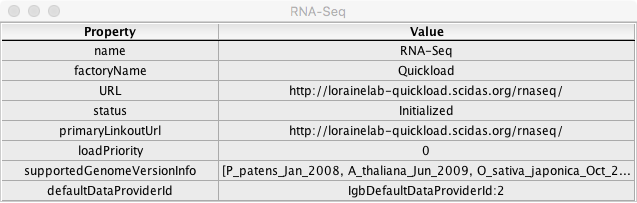
- Confirm that clicking on a [i] icon in the rightmost 'information' column of the Data Sources window above opens a window with fields and values similar to the example below:
- Confirm the following species / genome versions are available
- Arabidopsis thaliana / A_thaliana_Jun2009
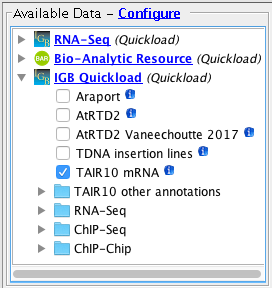
- Observation: Available Data Tree appears with several sites. (They may be different from those shown below.)
- Observation: Available Data Tree appears with several sites. (They may be different from those shown below.)
- Homo Sapiens/ H_sapiens_Dec_2013
- Available data should list some data sets. (They may vary from those shown below.)
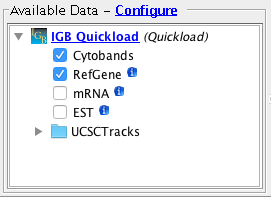
- Important: Confirm that for the human genome assembly, the Cytobands track is visible
- Available data should list some data sets. (They may vary from those shown below.)
- Arabidopsis thaliana / A_thaliana_Jun2009
...
- Load a few tracks from the UCSC DAS data source. Note: The UCSC DAS data source is not activated by default - you need to activate it in the Data Sources tab of the Preference window. Once this data source is active, it should be visible on Human/Mouse species.
- Adding/Removing/Editing DataProviders
- Add a quickload data provider and confirm it is working as expected
- Go to the Arabidopsis thaliana/A_thaliana_Jun_2009 genome version
- Navigate to the File->Preferences->Data Sources tab
- Click the "Add..." button
- Enter in a valid quickload url (e.g. http://www.igbquickload.org/abiotic)
- Enter a name for this quickload site and click the "Submit" button
- Observations:
- The quickload site shows up in the "Available Data" tree
- The data sets listed in the tree can be loaded
- Confirm restarting IGB does not cause the newly added site to be forgotten
- Ensure that the edit button in the Data Sources window opens a new window and allows a user to modify an app repository URL or to choose a local folder.
Ensure that data sources can be deleted using the Remove button in the Data Sources window
- Add a few "Secured" quickload sites and confirm they are working as expected
- Following the same steps listed above in the previous step, add the following two sites
- http://igbquickload.org/secureQuickloadTestSites/secureSiteTest
- username/password
- guest/guest
- username/password
- http://igbquickload.org/secureQuickloadTestSites/secureSiteTest2
- username/password
- guest2/guest2
- username/password
- http://igbquickload.org/secureQuickloadTestSites/secureSiteTest
- Navigate to the Arabidopsis thaliana Jun_2009 genome version
- Confirm the two sites are listed the "Available Data" tree and data can be loaded from each site
- Confirm you were not prompted for a password for any site more than a single time per session
- Confirm restarting IGB does not cause the newly added sites to be forgotten
- Confirm that if you checked the "Save Password" option you are not prompted for your password again on restarting IGB
- Following the same steps listed above in the previous step, add the following two sites
- Add a quickload data provider and confirm it is working as expected
...