General Function Checklist
This page tests local QuickLoads using relative file paths.
(To test absolute file paths see QuickLoad Saver release testing documentation)
Add files to your computer
- Download
...
- the following file:
...
- relative space.zip
- Extract it to where you would like your
...
- local QuickLoad to be stored. (Desktop, C drive, etc.)
...
- Files were extracted to local machine:
- mac
- linux
- windows
Add the QuickLoad to IGB
...
- Open the Data Sources tab
...
...
- in Preferences.
- Click
...
- Add…
...
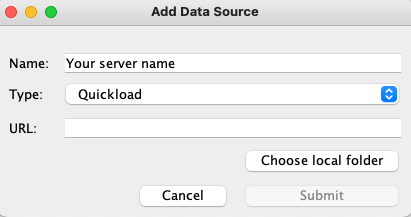
- to open the Add Data Source window.
...
- Click
...
- Choose local folder
...
- .
- Choose the unzipped QuickLoad folder
...
- “relative space”.
...
...
- Click Submit.
- The data on your QuickLoad is reachable by IGB and not highlighted in Red, Yellow, or Greyed out
...
- .
- mac
- linux
- windows
Access local QuickLoad data
...
- Close the Preferences window.
...
- Open the E_unicornis_Jul_2043 genome.
- The correct Species (E_unicornis) and Genome Version
...
- (Jul_2043) loaded.
- mac
- linux
- windows
The data should automatically load in IGB.
- Data loads automatically and appears the same as the above image
...
- .
- mac
- linux
- windows
- Your QuickLoad
...
Data source is present:
- appears in the
...
- Available Data section.
- mac
- linux
- windows
Zoom in at any location and select Load Sequence.to the following coordinates, then click Load Sequence: chrXVI:387,629-387,744
- The sequence data loads
...
- along the coordinates track.
- mac
- linux
- windows
...