General Function Checklist
...
Default Data Providers
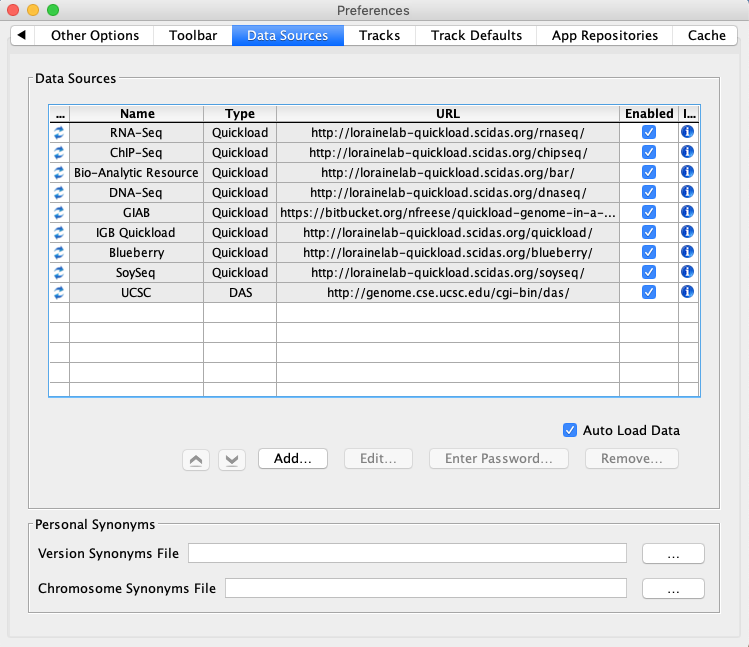
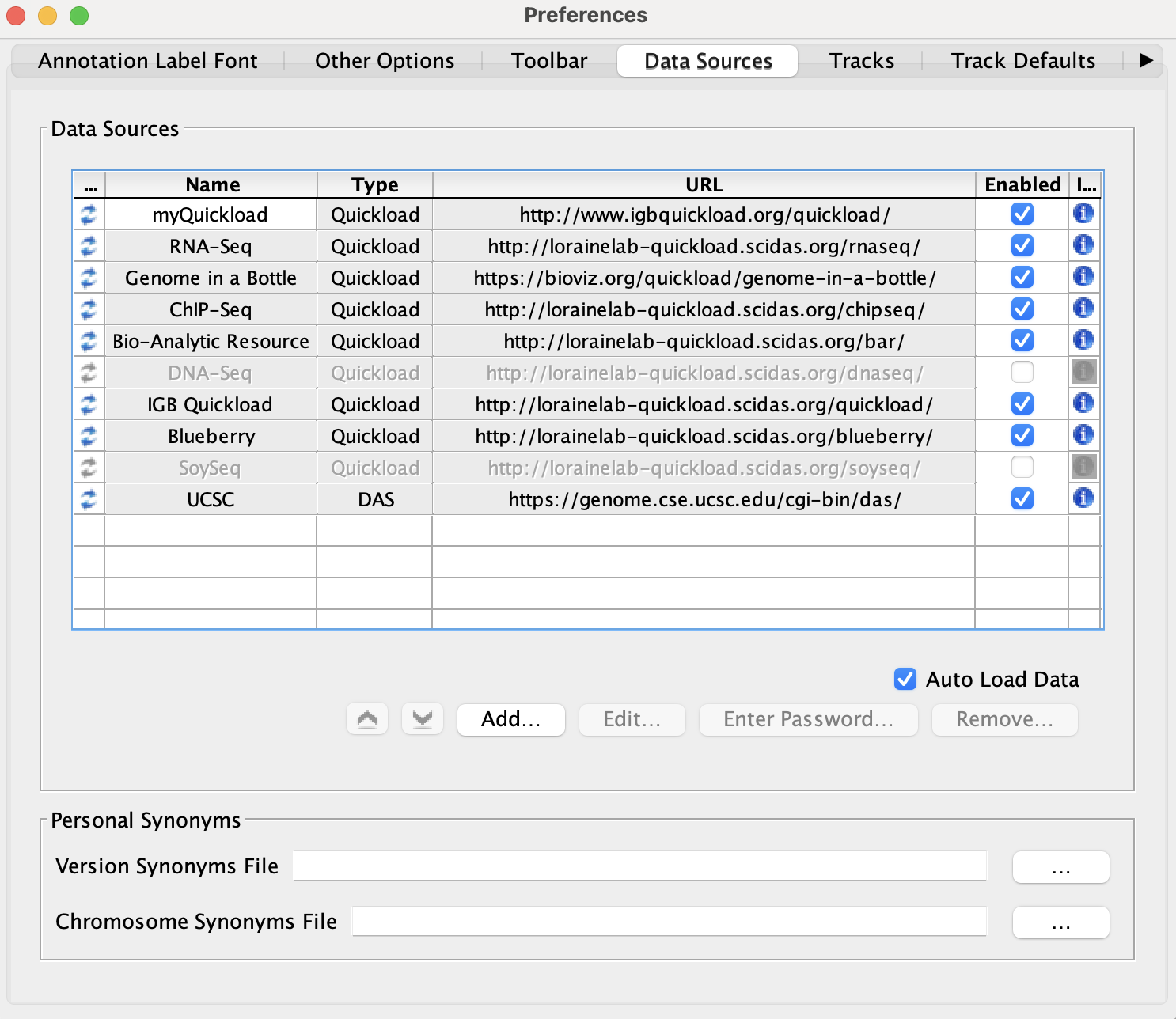
Select File -> Preferences -> Data Sources tab.
- Default data providers (e.g., RNA-Seq, ChIP-Seq, DNA-Seq, etc.) appear in the
...
- Data Sources table.
- mac
- linux
- windows
- Observation: The table lists several Quickload sites. (They may be different from those shown below.) If any are shown with a red background, that means that there is a problem with the site - IGB can't reach it. This is a bug and should be reported.
...
- None of the rows have a red or yellow background.
- mac
- linux
- windows
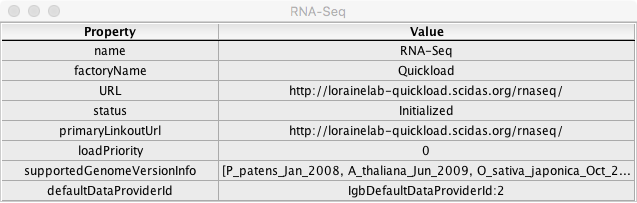
Click on the icon in the Information column for RNA-Seq.
- A window opened with fields and values similar to the example below:
- mac
- linux
- windows
...
- The data listed in the Value column are specific to the RNA-Seq data source.
- mac
- linux
- windows
Confirm that closing
Close this information window and opening another shows correctly updated information
- mac
- linux
- windows
...
Confirm that the information window functions for each data source type , then click on the icon for Genome in a Bottle.
- The data listed in the Value column updated appropriately and are specific to the Genome in a Bottle data source.
- mac
- linux
- windows
Confirm the following species / genome versions are available
- mac
- linux
- windows
...
...
Open the A_thaliana_
...
Jun_2009 genome.
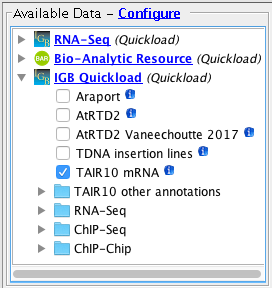
- Several Quickloads are present in the Available Data section (they may be different from those shown below).
...
- Homo Sapiens/
- mac
- linux
- windows
Open the H_sapiens_Dec_
...
2013 genome.
- Several Quickloads are present in the Available Data section (they may be different from those shown below).
...
- mac
- linux
- windows
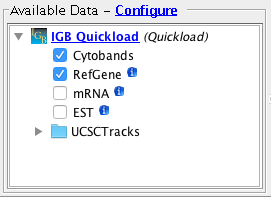
- Important:
...
- The Cytobands track is visible
- mac
- linux
- windows
...
- Load a bed file, a bam file, a bedgraph file, and some sequence data to ensure basic file parsing from the data providers is working as expected. Important: Load these by selecting the checkboxes in the Data Access folder hierarchy, not from URLs displayed in a Web browser. To get examples of each file type, go the the Arabidopsis thaliana June 2009 genome. Under RNA-Seq there are several folders labeled by experiment code and with a human-friendly name. Within those folders, there are folders labeled Reads, Graph, and Junctions. The Reads folder has bam files. The Graph folder has bedgraph files. The Junctions folder has bed files. Select one file from each folder, zoom in to show only several thousand bases, and click Load Data button at the top right of the IGB window. Click the Load Sequence button to load sequence and zoom in to make sure you can see individual letter (A, C, T, G) under the coordinate axis.
...
- Navigate to chr1:45,691,287-45,691,329
- Click Load Sequence.
- Individual base pairs (ACGT) are visible and colored consistently along the coordinates track.
- mac
- linux
- windows
...
- mac
- linux
- windows
...
Bedgraph File
- mac
- linux
- windows
...
Load Sequence
- mac
- linux
- windows
...
Adding/Removing/Editing Data Providers
- Select File -> Preferences -> Data Sources tab.
- Click Add...
- Name: myQuickload
- Type: Quickload
- URL: http://www.igbquickload.org/quickload/
- Click Submit.
- Close Preferences.
- myQuickload is present in the Available Data section
- mac
- linux
- windows
Adding/Removing/Editing DataProviders
Add a quickload data provider and confirm it is working as expected
- Go to the Arabidopsis thaliana/A_thaliana_Jun_2009 genome version
- Navigate to the File->Preferences->Data Sources tab
- Click the "Add..." button
- Enter in a valid quickload url (e.g. http://www.igbquickload.org/abiotic)
Enter a name for this quickload site and click the "Submit" buttonObservations:
...
...
- Close and re-open IGB.
- Select File -> Preferences -> Data Sources tab.
- myQuickload is still present in the Data Sources table.
- mac
- linux
- windows
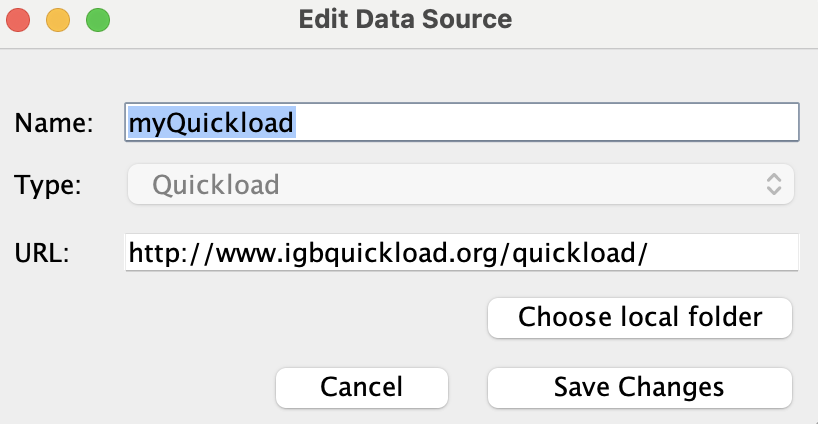
Select myQuickload and click Edit...
- The Edit Data Source window opened.
- mac
- linux
- windows
Confirm restarting IGB does not cause the newly added site to be forgotten
Click Choose local folder.
- The user's file chooser opens correctly.
- mac
- linux
- windows
Ensure that the edit button in the Data Sources window opens a new window and allows a user to modify the data source URL or to choose a local folder. Note that default data sources cannot be modified.
Change the Name of myQuickload to myQuickloadv2, then click Save Changes.
- The name of myQuickload correctly updated to myQuickloadv2 in both the Data Sources table and the Available Data section.
- mac
- linux
- windows
Ensure that data sources can be deleted using the Remove button in the Data Sources window
- Select myQuickloadv2.
- Click Remove...
- The myQuickloadv2 data source was removed from both the Data Sources table and the Available Data section.
- mac
- linux
- windows
Secured Data Providers
Add a few "Secured" quickload sites and confirm they are working as expected
...
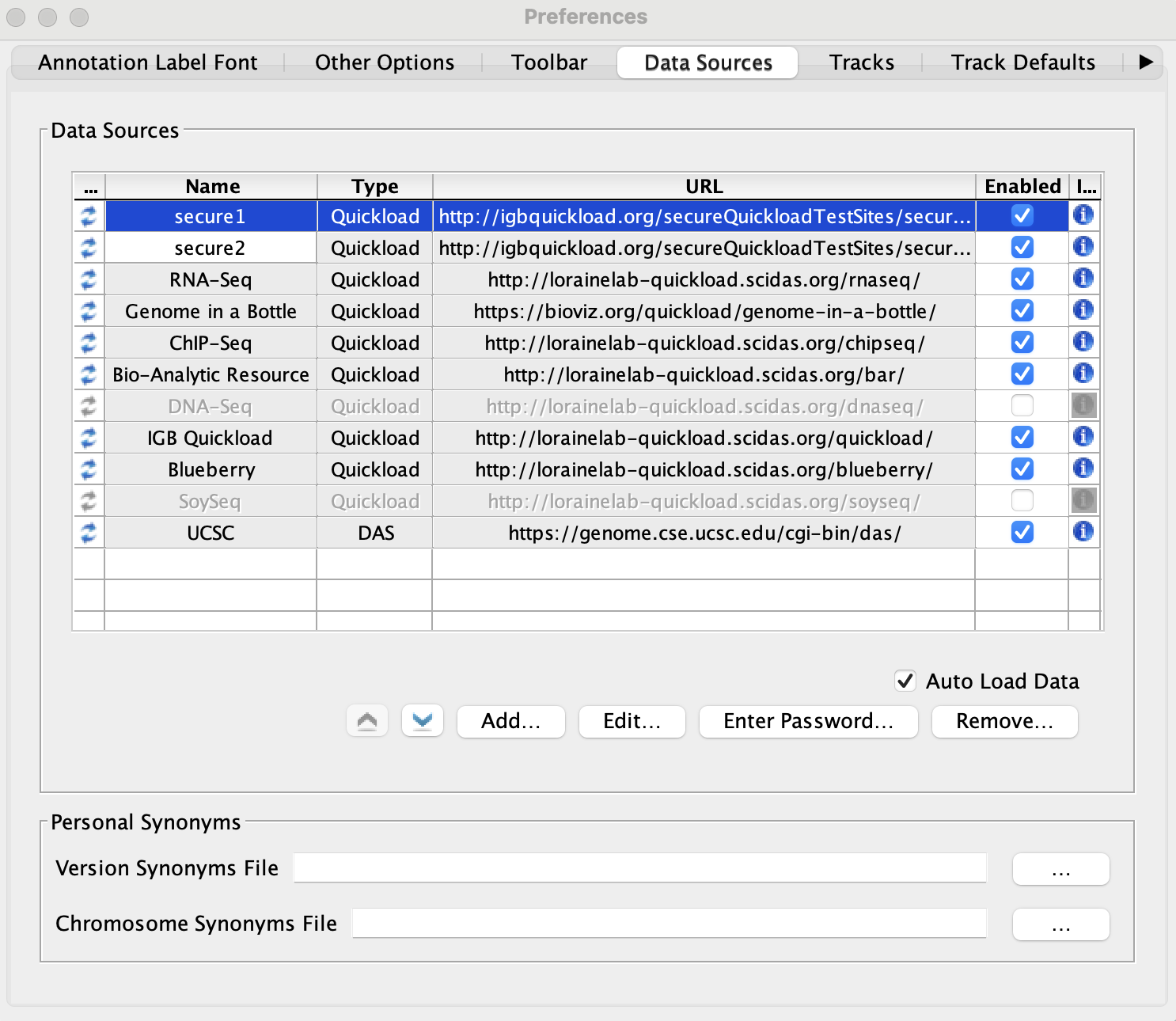
Click Add... and add the following two Quickload sites:
- Name: secure1
- Type: Quickload
- URL: http://igbquickload.org/secureQuickloadTestSites/secureSiteTest
username/password
- guest/guest
...
- Click Submit. A prompt should appear asking for a username and password:
- Username: guest
- Password: guest
- Make sure Save Password is checked, then click OK.
- Click Submit. A prompt should appear asking for a username and password:
- Name: secure2
- Type: Quickload
- URL: http://igbquickload.org/secureQuickloadTestSites/secureSiteTest2
username/password
- guest2/guest2
- Click Submit. A prompt should appear asking for a username and password:
- Username: guest2
- Password: guest2
- Make sure Save Password is checked, then click OK.
- Click Submit. A prompt should appear asking for a username and password:
- The two secured Quickloads are added to the Data Sources table and are not highlighted yellow or red.
- mac
- linux
- windows
Navigate to the Arabidopsis thaliana
Close Preferences and open the A_thaliana_Jun_2009 genome version.Confirm the
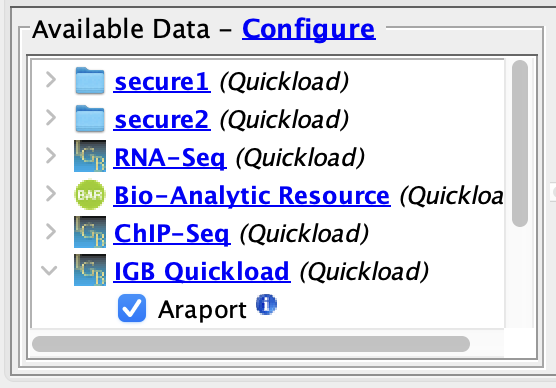
- The two
...
- secured Quickloads are listed in the
...
- Available Data section.
- mac
- linux
- windows
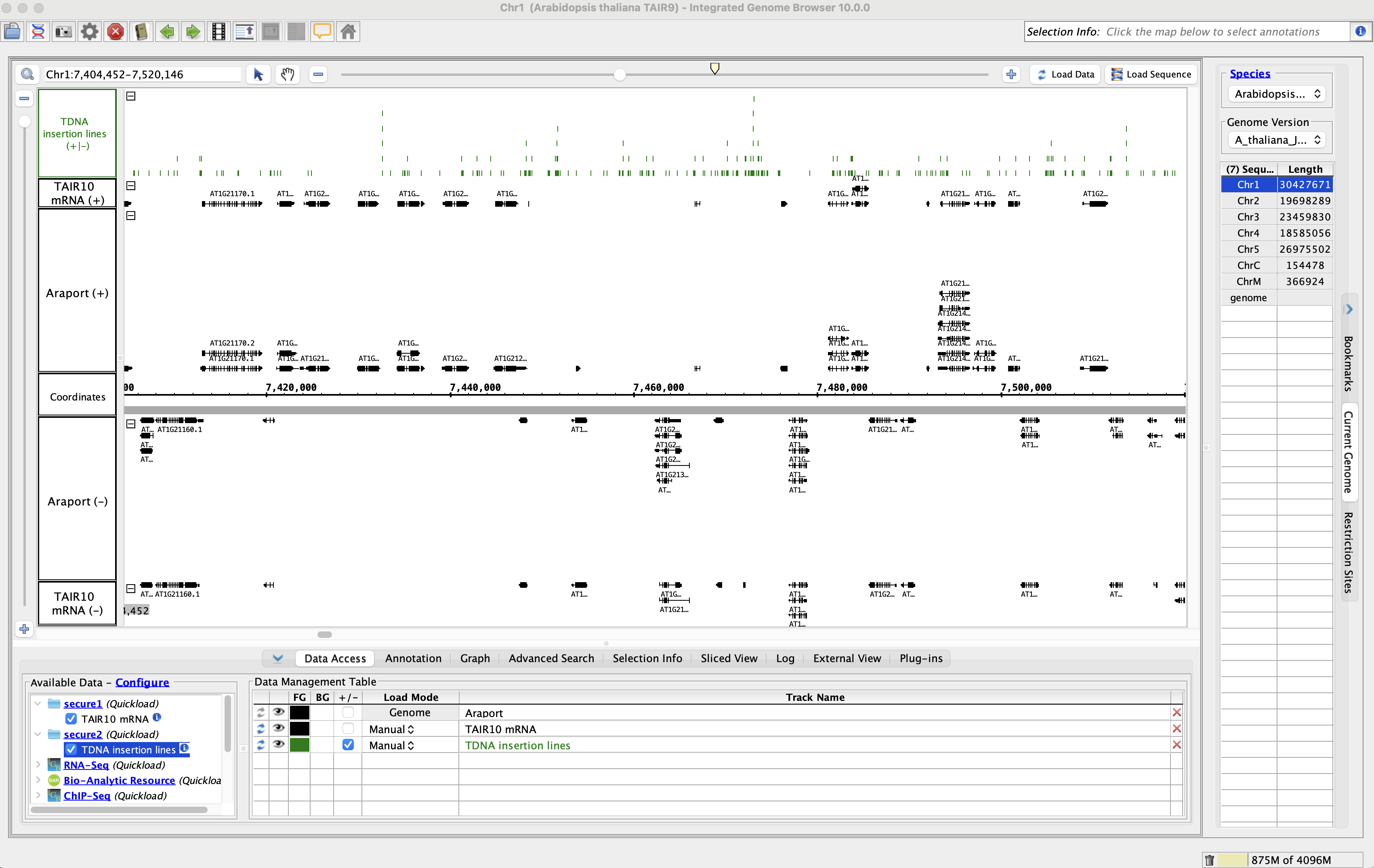
- Add the TAIR10 mRNA data from the secure1 Quickload.
- Add the TDNA insertion lines data from the secure2 Quickload.
- Go to Chr1:7,404,452-7,520,146
- Click Load Data.
- Data loaded from both of the secured Quickloads.
- mac
- linux
- windows
...
- You were not prompted for a password for
...
- either of the secured Quickloads.
- mac
- linux
- windows
Confirm restarting IGB does not cause the newly added sites to be forgotten
Restart IGB and open the A_thaliana_Jun_2009 genome.
- The two secured Quickloads are still listed in the Available Data section.
- mac
- linux
- windows
...
- You were not prompted for your password again
...
- for either of the secured Quickloads upon restarting IGB.
- mac
- linux
- windows