Convert UCSC track data hubs to IGB Quickloads
UCSC track data hubs are web-accessible directories of genomic data. As UCSC track data hubs have a similar organization to IGB Quickloads, we have created a website - translate.bioviz.org - that can create converted track data hub URLs that can be used in IGB as IGB Quickloads. This is accomplished through an API that dynamically converts the track data hub information to the IGB Quickload format that can be used by IGB.
More information about how UCSC track data hubs work can be found at: https://genome.ucsc.edu/goldenPath/help/hgTrackHubHelp.html
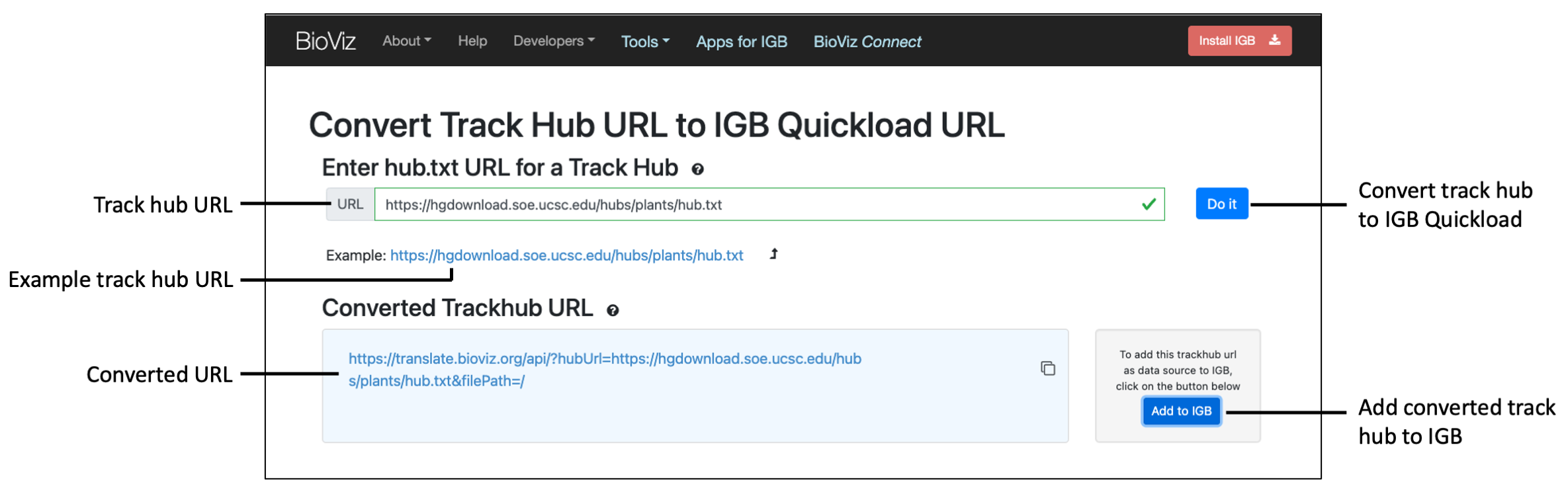
Steps to convert a UCSC track data hub URL to IGB Quickload URL
- In a web browser, navigate to https://translate.bioviz.org/
- In the URL text box, enter the track hub URL you would like to convert.
- For example: https://hgdownload.soe.ucsc.edu/hubs/plants/hub.txt
- Click the "Do it" button to convert the track data hub URL.
- If successful, the converted URL will appear in the blue box.
- Click the Add to IGB button to add the converted track data hub to IGB as a Quickload.
- Note: IGB 9.1.10 or later must be running.
View public UCSC track data hubs in IGB
Public Hubs are a collection of track data hubs that are of general interest to the research community and have been registered with UCSC. We utilized the UCSC API to create a website - bioviz.org/public-trackhubs.html - which populates the searchable list of Public Hubs. Public Hubs of interest can then be added to IGB by clicking the "+" icon.
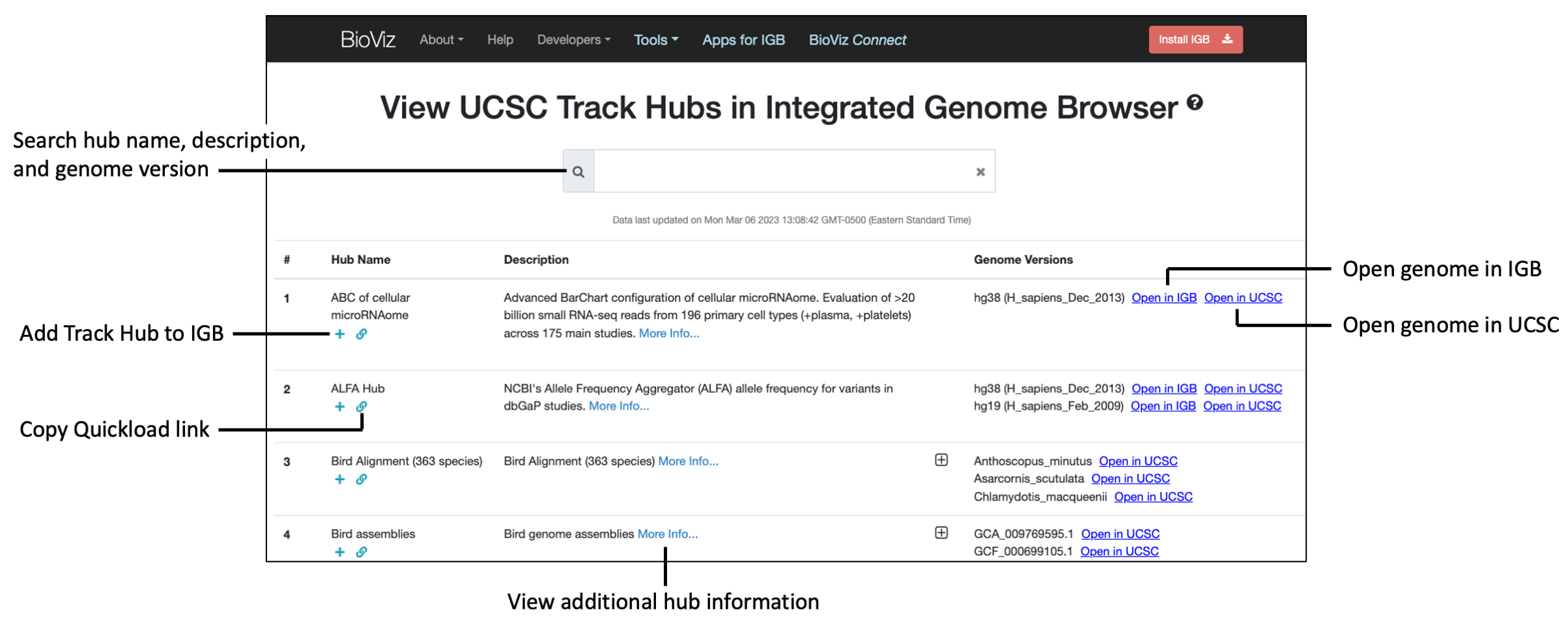
Steps to add a public UCSC track data hub to IGB
- In a web browser, navigate to https://bioviz.org/public-trackhubs.html
- Find a track data hub of interest using the Search bar or navigating through the list of public hubs.
- Click the "+" below the hub name to add the public hub to IGB as a Quickload.
- Note: IGB 9.1.10 or later must be running.
- If available, click the Open in IGB link to open the genome in IGB.