Use the Advanced Search tab to search sequence residues or annotations. Both search types support regular expressions and wild card characters.
Using advanced search you can
Search results will appear in the Advanced Search results table. Double-click a row in the table to view the result in the main IGB window.
If you search for sequence, IGB will also display color-coded bars in the coordinates track indicating the matched sequence.
The Advanced Search supports:
To find a feature or annotation by ID or name
Only data already loaded into the IGB viewer will be searched.
IDs or Names search will search IDs and names of annotations.
Keyword search, similar to ID or name search, will search annotation names, but it will also search other information associated with annotations, such as descriptions or other attributes. Choose Keyword to search descriptions, names, ids, and other attributes.
Results will appear in the Search Results table. Double-click a row in the table to zoom to that feature.
Advanced Search tabbed panel after searching the human genome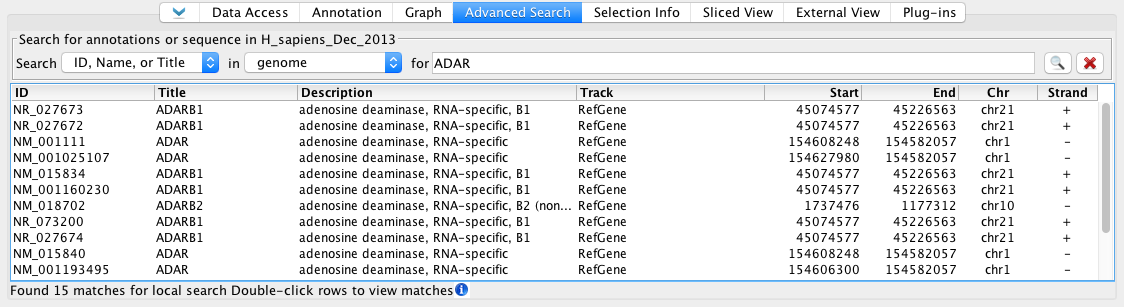
To find all instances of a sequence or regular expression
IGB displays matches in the results table and as colored bars underneath the coordinates axis. Matches on the minus strand appear in a slightly lower position.
Residue search results.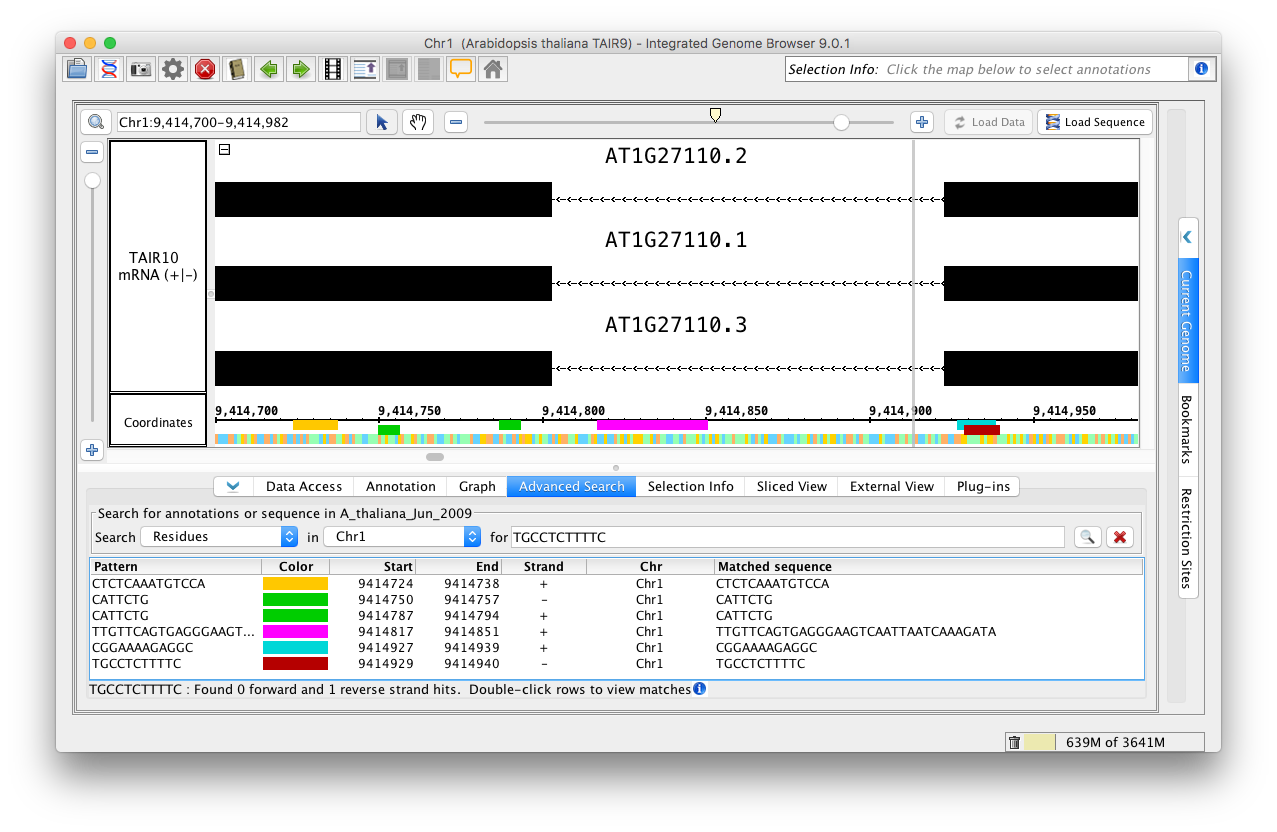
To search for multiple residues simultaneously enter residues in advanced search box separated with the pipe (|) symbol.
Example: atgttc|atggc
This will return a search for atgttc and atggc separately.
IGB searching supports regular expressions and wild cards. This is especially useful when searching for sequence motifs, such as transcription factor binding sites.
Searching by nucleotide symbols is available in IGB versions 9.1.12 and above.
Example queries:
Pattern | Represents | Example | Finds |
. | any single nucleotide | ACCT.T | ACCTTT, ACCTAT, ACCTGT, and ACCTCT (4 possibilities) |
.. | any two nucleotides | ACCT..T | ACCTAAT, ACCTATT, ACTAGT, Etc. (4 x 4 possibilities) |
[CG] | C or G | ACCT[CG]TC | ACCTCTC and ACCTGTC |
T{1,n} | 1 to n T's | ACGGT{1,3}C | ACGGTC, ACGGTTC, ACGGTTTC |
T* | Zero or more T's | ACGGT*C | ACGGC, ACGGTC, ACGGTTC, ACGGTTTC, ACGGTTTTTTTTTTTTTTTTTTTTTTTTTTTC, Etc. |
.*? | a string of any length containing any nucleotides | TCGGGGTTAA.*?CTGGACTC | Many possibilities. Because this allows for so many possibilities, it only recommended with a limited scope of search and/or with very specific (several specified base pairs) on both ends. |
.* | the longest possible string of any length containing any nucleotides | TCGGGGTTAA.*CTGGACTC | Differs from the search above in that the longest possible result(s) will be found. Bear in mind that the result returned from this search with depend on the scope of the search, ie how much of the genomic sequence has been loaded and is available for searching. |
| R | A or G | GCCR | GCCA, GCCG |
| Y | C or T | AGCY | AGCC, AGCT |
| S | G or C | AGCS | AGCG, AGCC |
| W | A or T | AGCW | AGCA, AGCT |
| K | G or T | AGCK | AGCG, AGCT |
| M | A or C | AGCM | AGCA, AGCC |
| B | C or G or T | AGCB | AGCC, AGCG, AGCT |
| D | A or G or T | AGCD | AGCA, AGCG, AGCT |
| H | A or C or T | AGCH | AGCA, AGCC, AGCT |
| V | A or C or G | AGCV | AGCA, AGCC, AGCG |
| N | any base i.e. A or G or T or C | AGCN | AGCA, AGCG, AGCT, AGCC |
| \QN\E | N | AGC\QNNN\E | AGCNNN |
More information about regular expressions is available from http://docs.oracle.com/javase/7/docs/index.html.