General Function Checklist
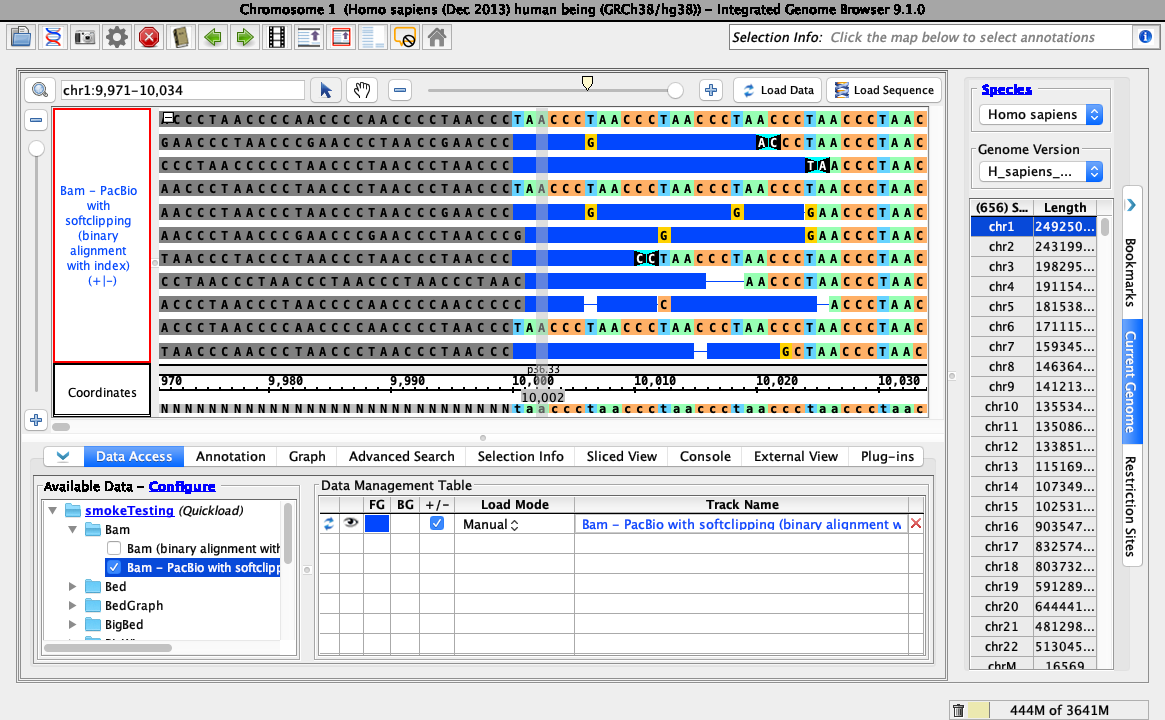
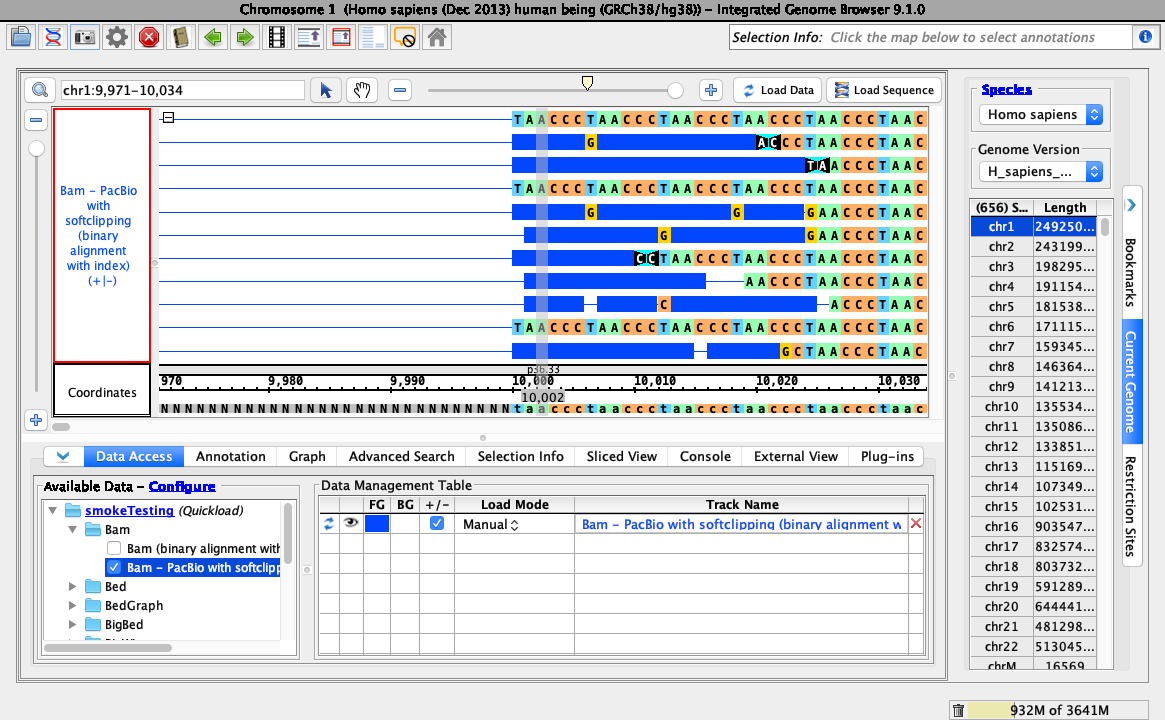
Objective: Validate that soft-clipping can be hidden and customized using the right-click menu of a bam track, as shown below:
...
- In the Data Sources tab in Preferences, add the smoke test Data Provider to IGB using the following URL: https://quickload-testing.s3.amazonaws.com/smokeTestingQuickload/
- Open
...
- the H_Sapiens_Dec_2013 genome.
- In the Available Data panel, add the Bam - PacBio with
...
- softclipping data from your newly added Quickload.
- Navigate to: chr1:9,971-10,034
- Click Load Sequence
...
- to load sequence data,
...
- then click Load Data to load data into it.
Ensure that loading
- Loading data looks like this (stack height set at 10, reads may be organized randomly):
- mac
- linux
- windows
Validate that the '
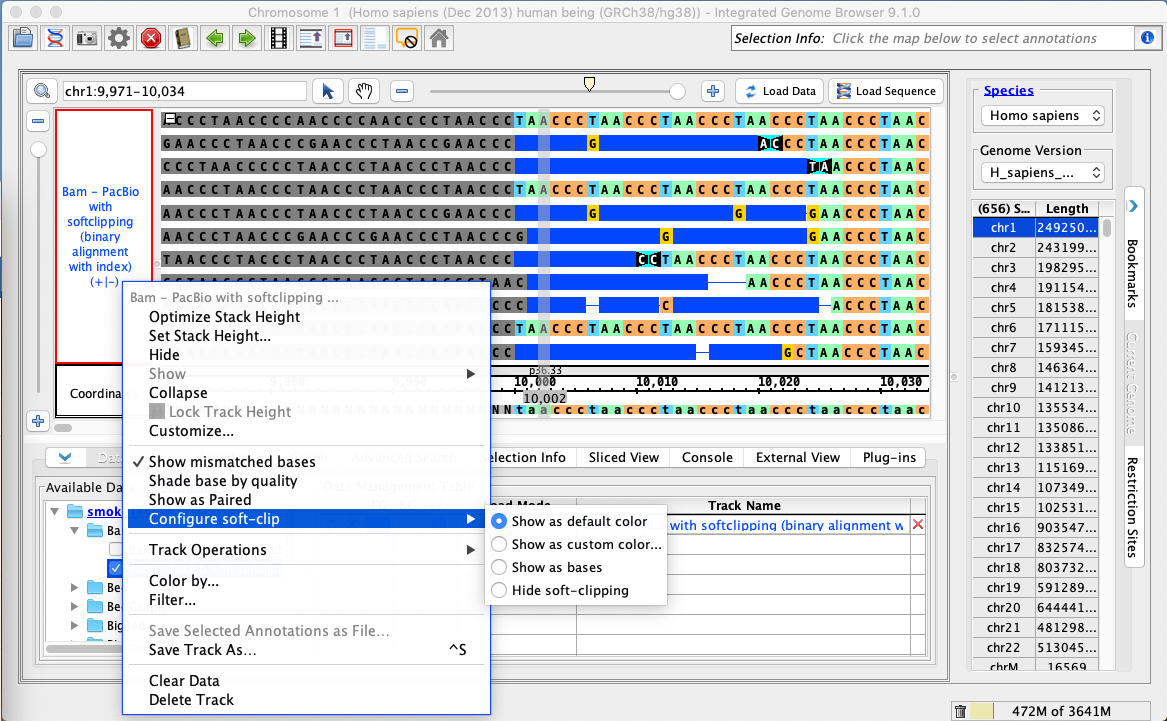
- The Configure soft-clip
...
- submenu of the bam track right-click menu has
...
- Show as default color
...
- selected.
- mac
- linux
- windows
- Right-click on the bam track and select:
...
- Configure soft-clip -> Show as custom color
...
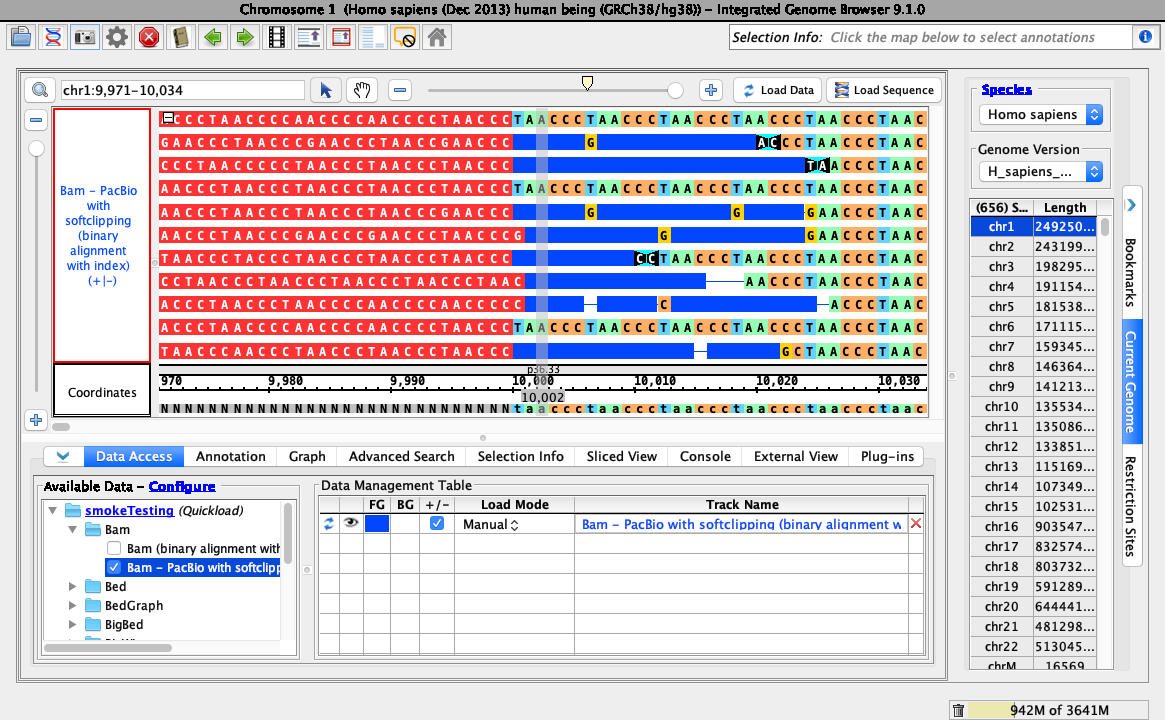
A color picker should appear. Select a color (the example below selected red).
...
- .
- Choose a red color from the color picker.
- Your IGB looks like the above image.
- mac
- linux
- windows
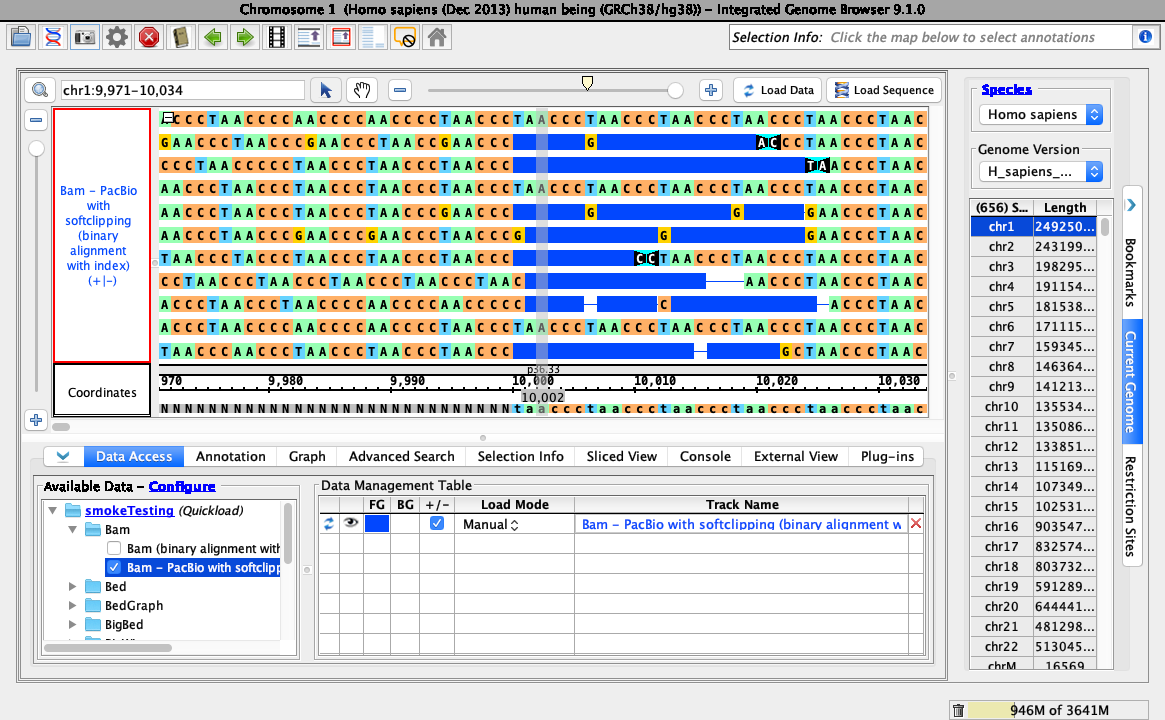
Right-click on the bam track and select:
Configure soft-clip -> Show as basesEnsure your view is as below:.
- Your IGB looks like the above image.
- mac
- linux
- windows
Right-click on the bam track and select:
Configure soft-clip -> Hide softclippingEnsure your view is as below: .
- Your IGB looks like the above image.
- mac
- linux
- windows
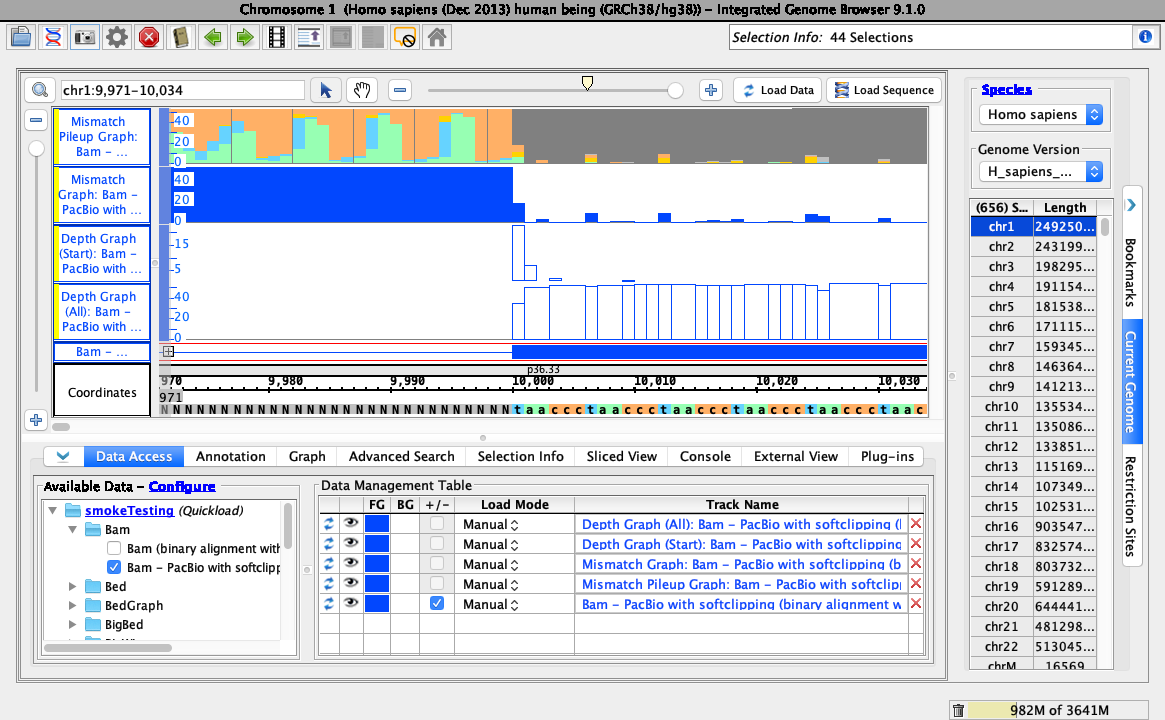
To test track operations are working correctly for soft-clipped data, right right-click on the bam track and select the following:
- Track Operations > Depth Graph (All)
- Track Operations > Depth Graph (Start)
- Track Operations > Mismatch Graph
- Track Operations > Mismatch Pileup Graph
- Collapse
Ensure your view is as below:
- Your IGB looks like the above image.
- mac
- linux
- windows