| Table of Contents |
|---|
Introduction
As you learn to use IGB, you'll find it offers many features that make it one of the best tools available for visualization and exploration of genomic data sets.
If you are new to IGB, the following six step use this Quick Start Guide will help you get started using IGB.
| Table of Contents |
|---|
Step 1: Get and start IGB
- Go to biovizBioViz.org/igb/download.html IGB Download PageDownload and launch IGB. and click the Download button
- Select and download the installer for your platform
If you have trouble starting IGB, see visit the Troubleshooting page or feel free to contact us. help page on BioViz.org for assistance.
Step 2: Choose species and genome version
...
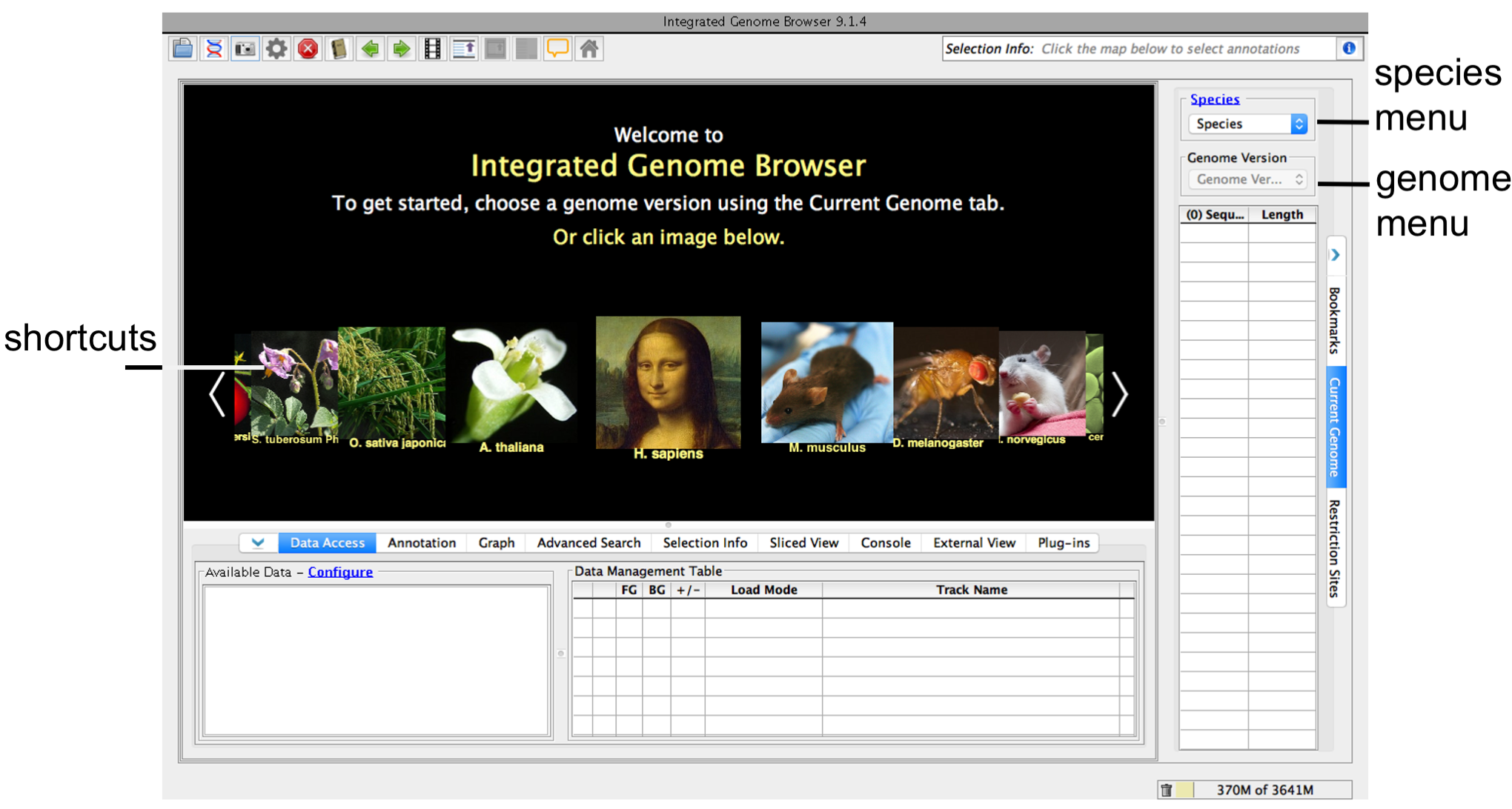
- Click a shortcut image (loads most recent genome for selected species)
or
- Choose Species and Genome Version using the Current Genome tab.
IGB start screen
...
If your species or genome version is not listed, you can view
...
See also
it if you have a fasta or 2bit file with sequence data. See Custom Genomes
Step 3. Open data sets
You can open Open data sets from remote data sources (Data Access tab) or by opening local files.
To open remote data setsa data set from an IGB Quickload data source (see About IGB Quickload)
- Select Data Access
- Select data sets in the Available Data sub-panel section
To open local files on your computer
- Select File > Open File... or File > Open URL...
or
...
- Enter file
...
- name or URL
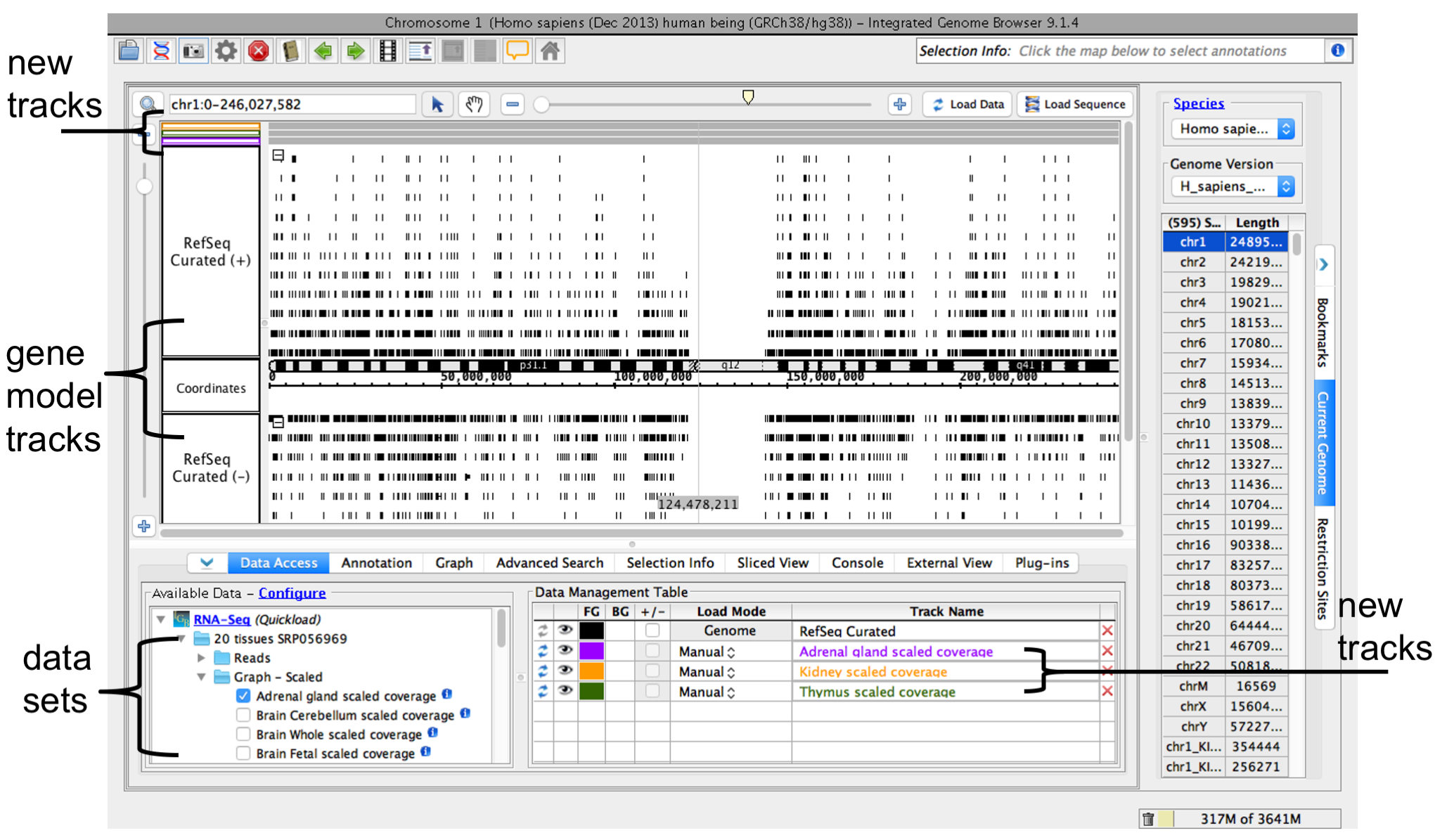
When you select a data sets or filesa file, IGB adds a new empty track to the main view and lists it in the Data Management table. Empty regions in the new track that do not have data loaded are gray.
IGB window after opening a data set
Genome reference sequence
If you would like to open a sequence file (fasta or 2bit format) to use as the reference sequence
- Select File > Open Reference Sequence...
human genome RNA-Seq coverage graphs from Adrenal Gland, Kidney, and Thymus data sets
Step 4: Zoom in
Because many data files contain too much data to view all at once, IGB does not start loading load data into the viewer until you click the Load Data button.
Before loading data, zoom in to a region you want to view.
To zoom in on a region
- To focus zooming, click Click a location in the main view
- To zoom in, drag Drag the horizontal zoom slider to the rightor use plus and minus buttons
IGB horizontal zooming controls
Other ways to zoom
Other ways to zoom include
- Search for a gene using the search boxby name or keyword (For example, TBATA or thymus)
- Double-click a feature an exon or gene model to zoom in on it
- Click-drag over the coordinate axis to zoom in on a region
- Click the plus and minus buttons next to zoom sliders
See also:
Step 5: Load data
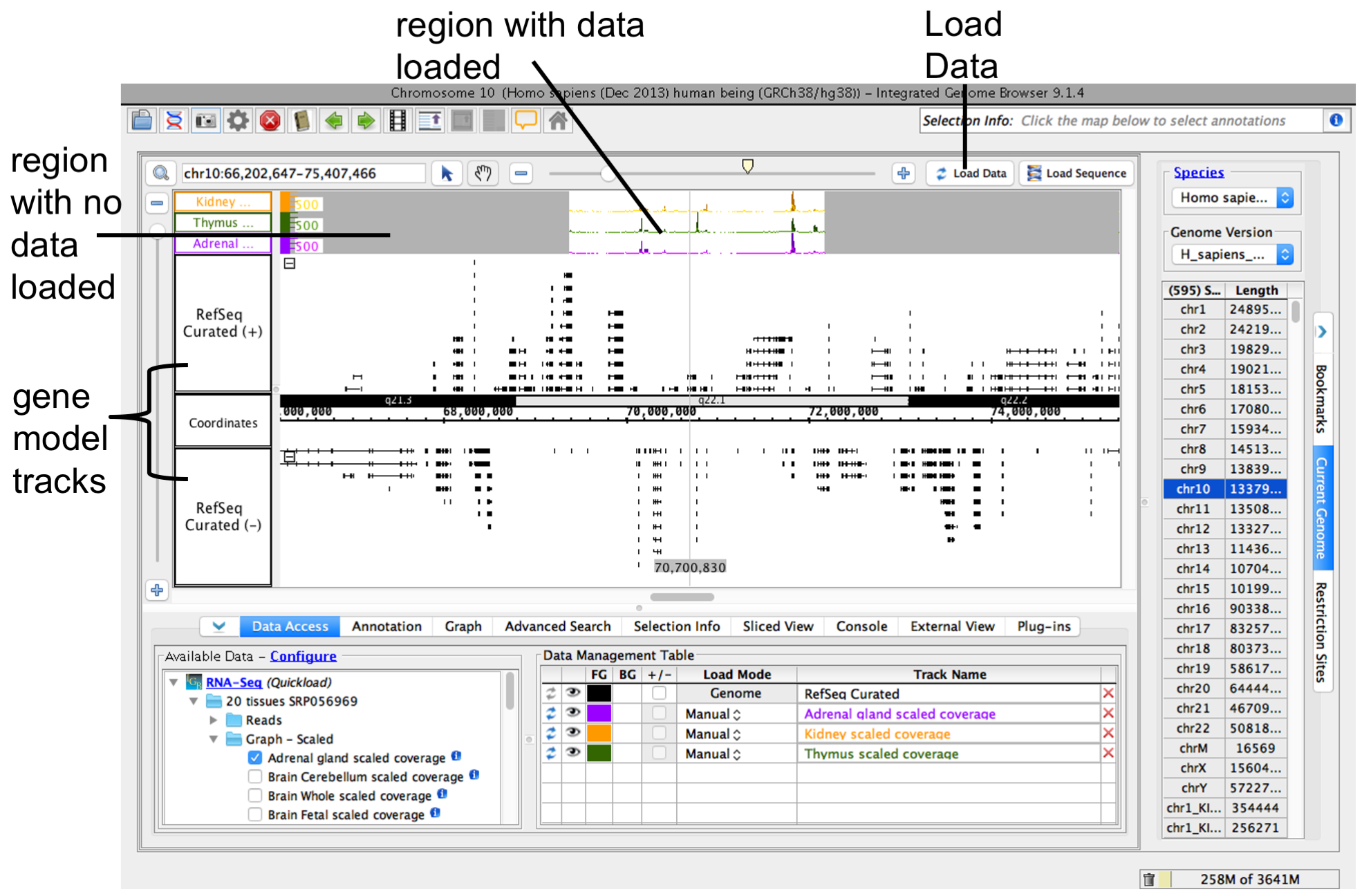
To load data for the currently visible region, click Load Data button. Regions with loaded data show the selected background color; areas without loaded data appear darker.
IGB after loading data
See also:
...
You can reorder the tracks by dragging the Track Label into the position you want (the Data Management Table reflect changes).
The Annotation and Graph tab will allow you To change style elements of a track, click the track label and use the Annotation or Graph tab to change to change color, track height, sizeannotation label, amount of data shown as well as other various functions(stack height), and other options.
See also: