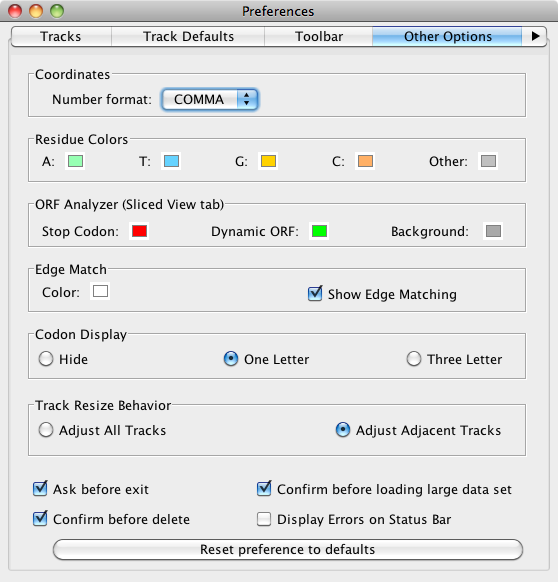
This tab contains several options addressing the color/appearance of non-track elements in the image, as well as some other miscellaneous options.
Axis
The options for axis appearance are here. You can change the foreground and background colors to suit your needs. Changes to the colors will not appear dynamically, but will take place when the Preferences window is closed.
...
| Table of Contents |
|---|
Introduction
Select File > Preferences > Other Options to configure the following:
- Coordinates track number format
- Sequence base colors
- ORF analyzer
- Edge matching
- Gene model translations
- Track resizing
- IGB exit and data set loading behaviors
Preferences Other options
Coordinates
These are options for changing number appearance. FULL shows the number with no thousands separators and the unit 'kb'. COMMA shows the numbers with commas at every 10^3 (American U.S. style) and with no unit designations. This is the default appearance. ABBREV will show the numbers as abbreviated as is reasonable, with unit designations, i.e. zoomed all the way . For example, when zoomed out, numbers are appear as 10M, 20M etc. Zoomed in a bit, numbers are shown as 19,500k, 20,500k. Zoomed all the way in, numbers are shown fully. Changes made here appear dynamically in your image.Changes made here to Number Format take effect immediately.
Residue Colors
IGB displays letters representing DNA bases (residues) when zoomed-in. Residues are shown in alignment tracks, sequence tracks, and in underneath the coordinate axis.
By default, IGB uses cool colors for A and T residues and warmer colors for G and C residues. Other refers to residues that are either 'N', 'X' or '-' (missing) and can be set to a bright color if you are looking for deletions in short read sequences; otherwise you may wish to leave it as the neutral gray.
Click a color swatch to change base colors.
ORF (Open Reading
...
Frame) Analyzer
This set of preferences changes the color of the stop codons and the ORFs These preferences affect colors used to indicate stop codons, the ORFs and the background of the ORF tracks that are shown in the Sliced View tab. After setting new color options, uncheck and recheck the Analyze ORFs option in the Sliced View tab to show the new color scheme.
Change Residue Colors
IGB shows each residue with an individual color highlight at the fully zoomed in level. At less zoom, where the letter cannot be fully shown, IGB will still show the color for each residue. In this section, you can change what color represents each residue individually; Other refers to residues that are either 'N' or '-' (missing). If you do NOT wish for individual residue highlights, set the colors to white. Changes made in this section will be applied to the imageafter the Preferences window is closed.
...
.
Edge Match
Edge matching highlights features that have identical boundaries. Edge matching becomes active when you click items in the display.
Click the swatch to change the edge match color. The selected color applies all annotation tracks.
| Tip |
|---|
Choose a color that has good contrast between foreground and background colors |
Codon Display
The Codon Display is activated when the sequence of a gene model is loaded and the user zooms in close enough to see the sequence. The codon translation appears within the gene model. User can set the amino acids to appear in one letter format (default), three letter format or be hidden. The colors is based on the chosen foreground color for the track.
| Info |
|---|
Codon display is active only for tracks loaded from 14-column BED detail files. |
Track Resize Behavior
Track Resize Behavior is how IGB adjusts other tracks when a specific track height is changed. Adjust All Tracks means that as a single track is increased in height, all other tracks are reduced by a commensurate amount. Adjust Adjacent Tracks (default) means that all added or subtracted height is removed/added to the bordering track, leaving the rest of the display unaffected.
Data loading and exit behaviors
Confirmations
IGB warns you when you are about to exit the program , and before you delete a file or files, and before you load very large files, >100MB. Each of these warnings has the option Do to not show be shown again. If you decide that you do want the warnings to appear, you can reset your choice from this pane. Check (or uncheck) Ask before exiting, Confirm before delete and/or Confirm before load.
Sometimes, the zoom stripe obscures sequence that you are interested in, or is otherwise distracting. You can hide the zoom stripe entirely by unchecking Keep zoom stripe in view.
If many changes have been made throughout the IGB interface, and you would like to start over (new user for instance) or if IGB is behaving 'oddly', you can Reset all preferences to defaults. Be careful, as this WILL reset ALL preferences by deleting the preferences file. You cannot undo this function!
To turn off these warnings or reset your previous choice, check (or uncheck) these options.
Reset all preferences to defaults
Click Reset preferences to defaults to reset all the preferences to the IGB defaults. This is sometimes useful if you need to troubleshoot IGB.
| Warning |
|---|
Be careful, as this action will reset ALL preferences and will also cause IGB to forget the location you were last viewing. |