...
| Table of Contents |
|---|
Probe set alignments - link.psl files
IGB can be used to visualize data from Affymetrix showing the locations of GeneChip Expression Array probe sets and target sequences aligned to a genome.
In most Affymetrix arrays, probes are grouped conceptually into probe sets, groups of probes that are expected to measure expression for individual known or computationally-deduced mRNA molecules.
These target sequences may be identical to known mRNA sequences in GenBank, or they may have been produced computationally by merging ESTs or mRNA sequences into a single sequence, sometimes called a "consensus" sequence.
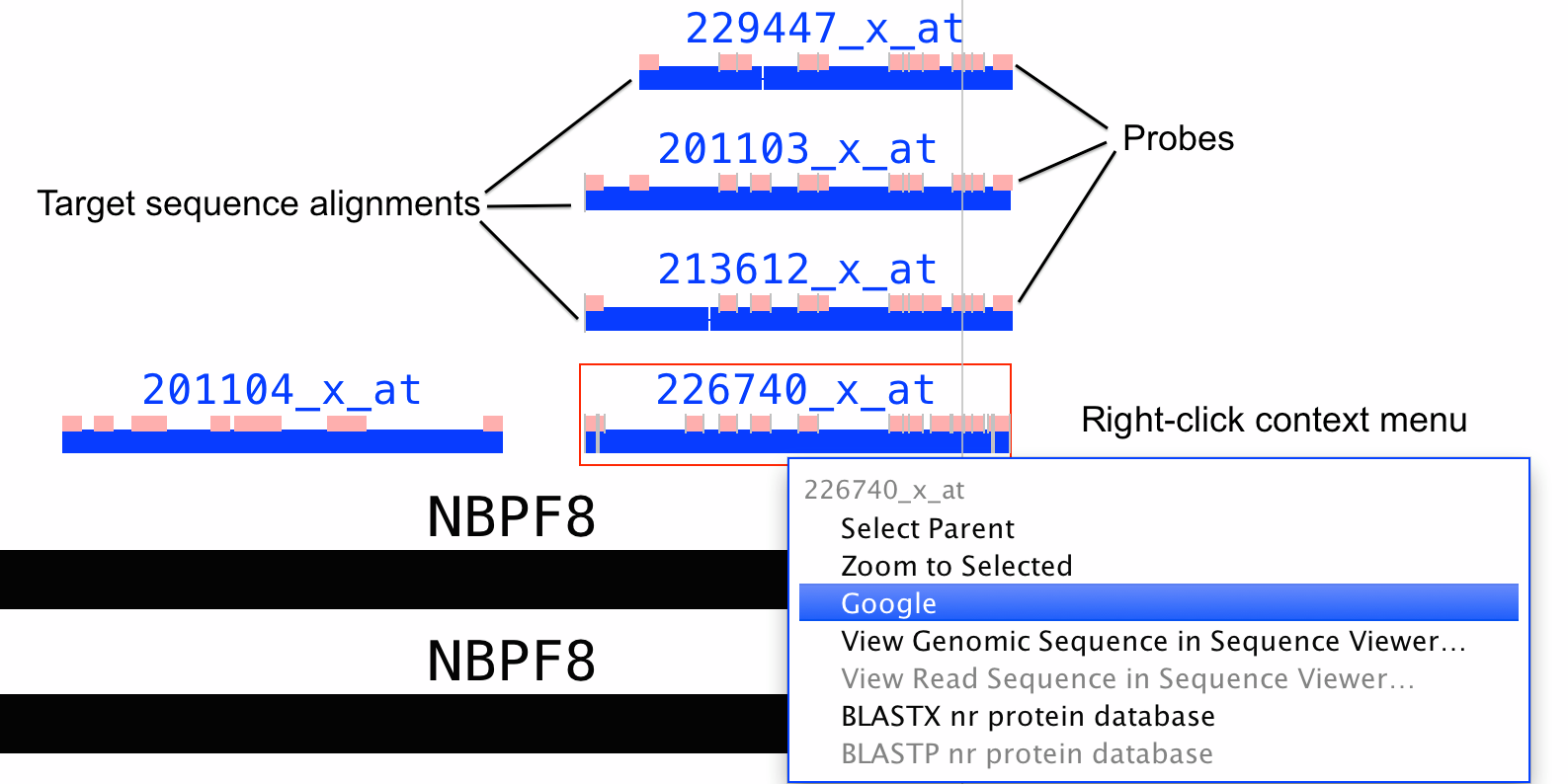
IGB can be used to visualize the location of design sequences and probes within the genomic sequence. Probe set alignments are available under the IGB QuickLoad DAS2 data source in IGB.display Affymetrix probe sets aligned onto a reference genome - it can show probe set design sequences aligned onto a genome with the locations of the probes indicated as blocks.
When IGB was first developed at Affymetrix, the company distributed probe set alignment files for its catalog 3' IVT arrays. In recent years, however, they've stopped updating these files. So for some genomes, the alignment files you can find on the Affymetrix Web site reference obsolete reference genomes. If you need to work with more up-to-date genomes, we recommend you create your own alignment files or request them from the IGB team. For some genomes, we've added probe set alignments to the main IGBQuickLoad.org site. Mainly we've done this at the request of individual researchers, and so if you would like to request an array, let us know. If probe set alignments are available from our site, you'll typically find them in a folder named Affymetrix under the Data Sources section of the Data Access panel.
You can also make your own probe set alignment files using blat, tabix, and a python script we wrote. For more information, see this Bitbucket repository.
About probe set alignment visualizations
Depending on when the arrays were designed, Affymetrix typically used expressed sequences from GenBank to select probes for probe sets - these expressed sequences were sometimes called "exemplar" or "consensus" sequences. They then selected individual probes from regions near the 3' end of the expressed sequence. Affymetrix (as of 2014) distributes probe and target sequences on their Web site, where "target sequences" contain the 3' end regions from which the probes were selected.
Probe set visualizations in IGB show the alignments of target, exemplar, or consensus sequences onto the genome. They also show the locations of probes that were selected from the design sequence. See the preceding figure for an example.
Because probes were selected from the expressed sequences, sometimes a probe will be shown as split across an intron. Also, sometimes probes overlap. And sometimes probes may be missing. If the target sequence contains a region that can't align onto the reference, and if this unaligned sequence contains a probe, then that probe will not be shown.
If you have questions about what you see in a probe set alignment, let us know.
Why this is useful
Often multiple, seemingly redundant probe sets interrogate one gene. This situation mainly arises when a gene has multiple, alternative three-prime ends due to alternative splicing or alternative termination sites. If an experiment identifies genes where redundant probe sets are differentially expressed with different fold-changes or in opposite directions, this can indicate that the treatment affects splicing as well as the overall abundance of RNAs arising from the gene.
Thus if you observe redundant probe sets that give different or contradictory results, it's a good idea to view them in IGB and compare their alignment to annotated genes and transcripts.